Genomic information at the service of human health and its environment
Bioinformatics

Explore our bioinformatic reports below.
Discover our report (de-novo assembly)
Discover our report (differential gene expression)
Interactive report
-
Interactive graphs and tables
-
Enriched data information
-
Data exploration directly from the report
-
Possibility to zoom in and to filter the data
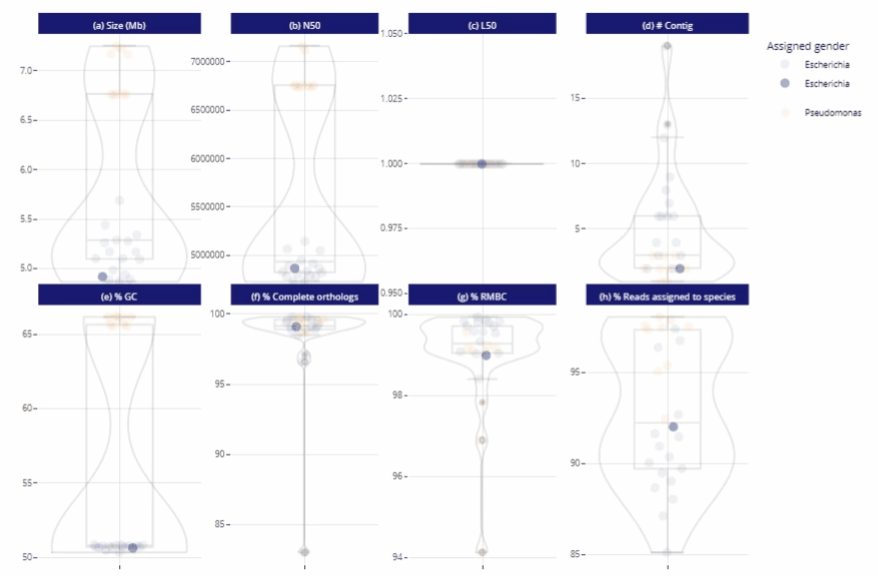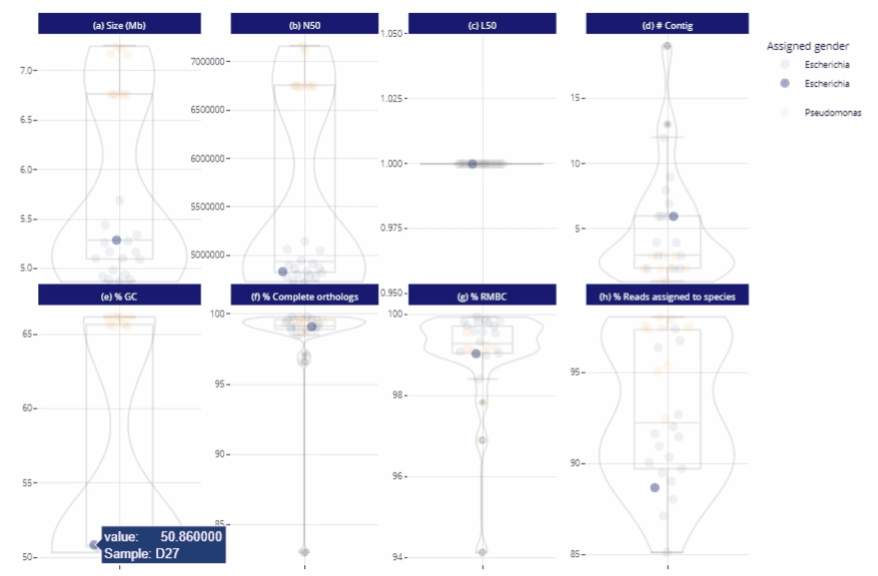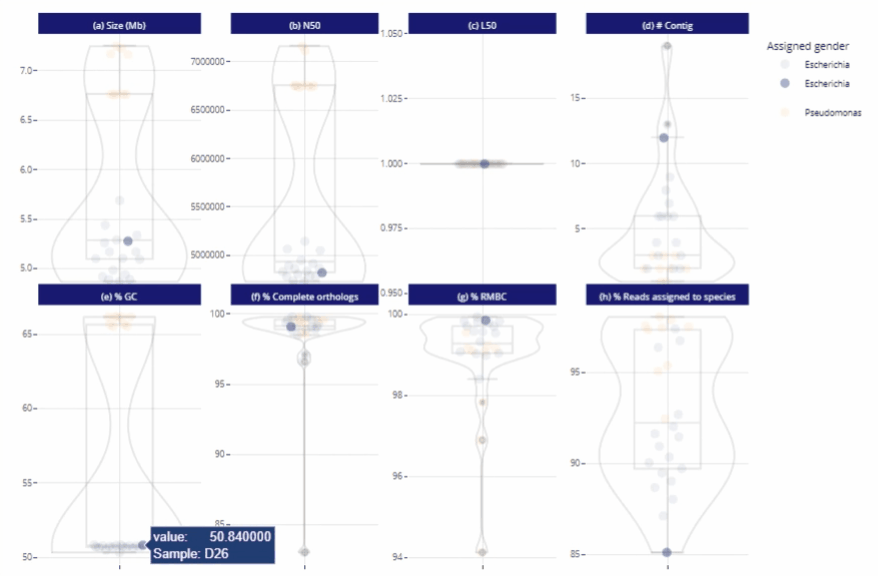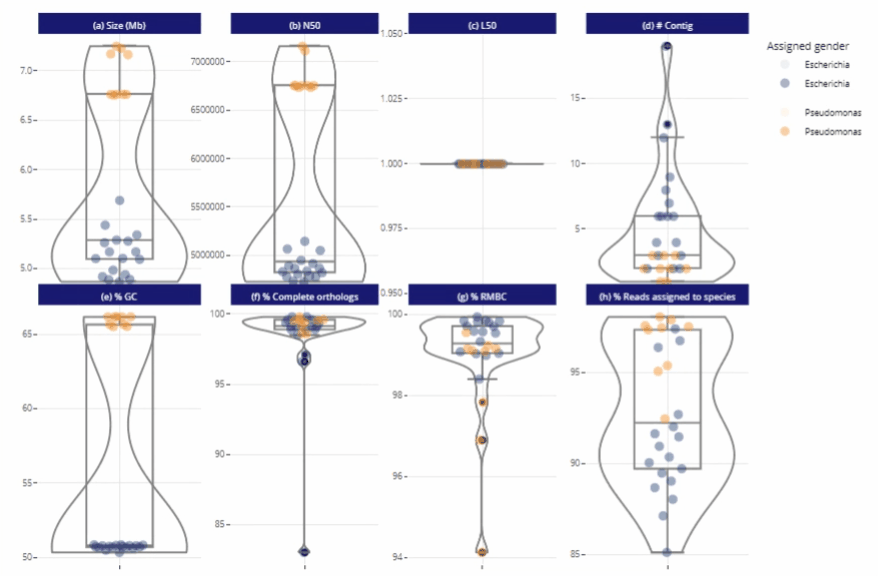
Different reading levels
FROM NEOPHYTE TO BIOINFO EXPERT
-
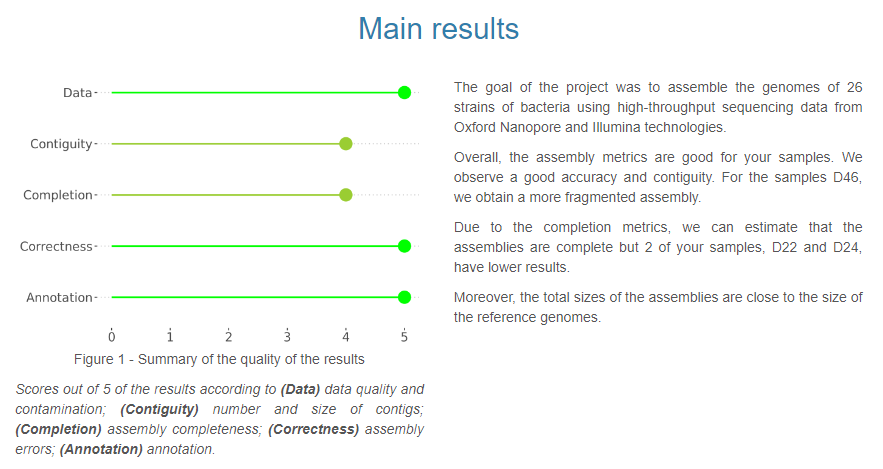
First page with both graphical and written summary
-
Each analysis step individually detailed
-
Visual highlighting of key results

Suitable for any size of project
-
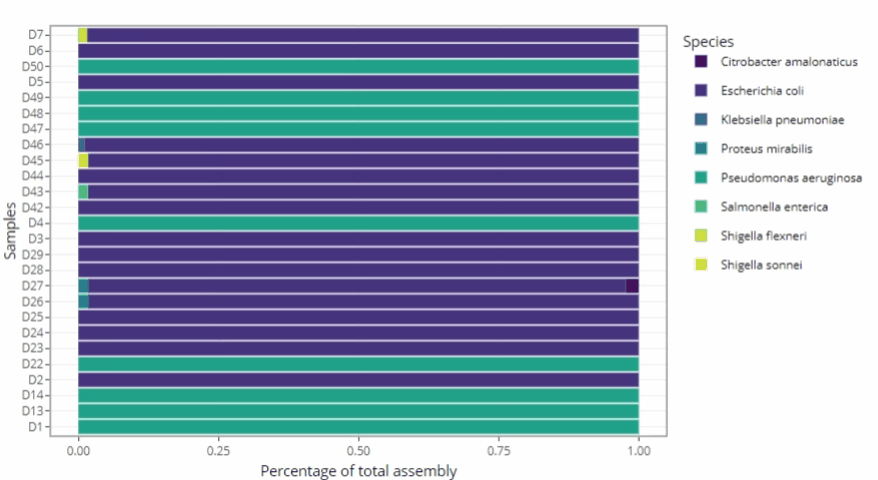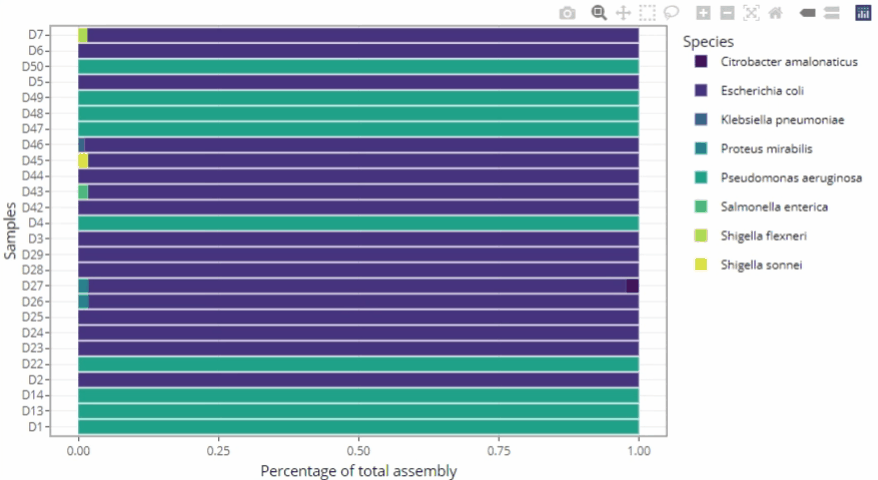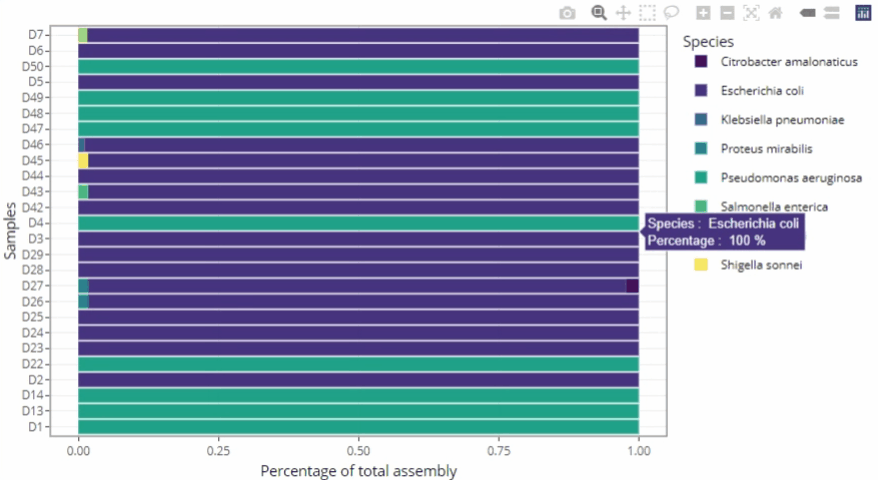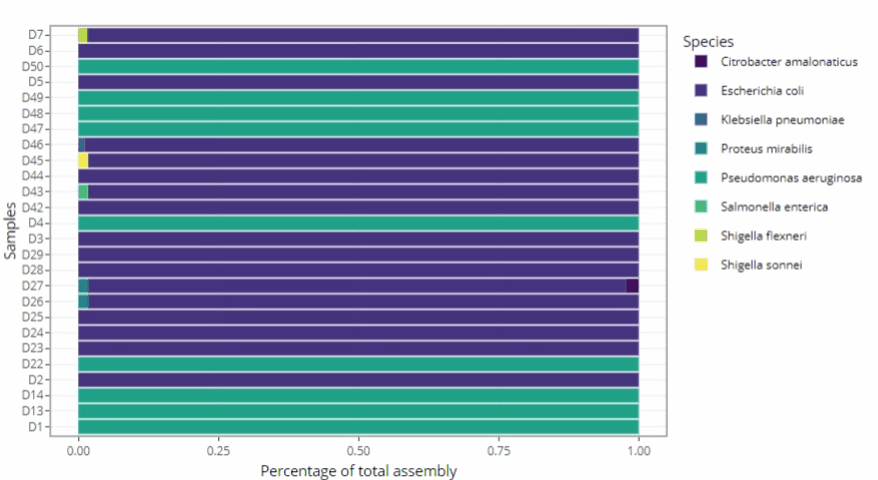
Graphical representations automatically adjust to the number of samples
-
Remains clear and readable for a single sample as well as for hundreds
Ready-to-publish methodology
-
Detailed material and methods section
-
Transparency on the tools used
-
References and sources already formatted for publication
One of the biggest challenges in genomics is being able to connect a phenotype to the available genetic information. Mutation detection and whole-genome comparison techniques can go beyond the simple observation of a genome, in order to make an evolution in your researches.
Applications of genome comparison and mutation detection
Whole-genome comparisons and mutation detection are deployed to identify genes and mutations of interest with regard to a molecular or cellular phenotype (antibiotic resistance, enzyme activity, etc.).
These two approaches make it possible to set up new genomic selection criteria in agriculture, identify drug targets, select specific strains, and discover novel biomarkers, for example.
GenoScreen offer - Mutation detection
Mutations of interest (SNPs or indels) are detected by comparing similar sequences (e.g. genomes/transcriptomes from two members of the same species).
This comparison involves mapping the sequences on a reference sequence. The segment of the reference genome covered must be similar for the two compared sequences.
The GenoScreen's assets
- Specific processing of sequencing information in order to improve their quality and clarity.
- Detailed annotation of genes and sequences of interest.
- Precise algorithms for alignment with reference sequences.
- Whole-genome comparisons.
GenoScreen develops a broad range of bioinformatics services - notably those designed to help interpret data generated by NGS. Our algorithms provide clear, precise answers to all types of genome-related issues.
Exclusive tools for interpretation and analysis
Our bioinformatics services are permanently powered by the results of our R&D programs. GenoScreen detains tools for post-processing raw sequencing data - guaranteeing the highest possible quality of information.
Many specific algorithms have been developed in collaboration with our genomics experts in order to provide a faster results interpretation and also a better understanding and a denser scientific information.
Gene expression is a major field of investigation with the goal of a better understanding of the mechanisms of action that underlie a cell’s behavior. GenoScreen offers a comprehensive range of differential genomics analyses, so that you can identify biomarkers of interest.
Applications of differencial genomics
RNA-Seq technologies provide the differential expression analysis of an entire genome in an organism and the comparison of the level of gene expression between two conditions, for the research of biomarkers of interest :
- Comparisons of healthy and diseased tissues.
- Comparisons of cell behavior in two different environments (as a function of nutrition, stress, pollution, etc.)
- Comparisons of cell behavior at different stages of biological development.
This exhaustive approach has several advantages:
- No requirement for prior knowledge of the organism being studied.
- Whole-transcriptome analysis, hypothesis-free on the genes of interest.
- The ability to use sequencing data to generate a transcriptome de novo.
- Analysis of natural antisense transcripts (NATs).
GenoScreen - Differencial genomics services provided
Our teams offer comprehensive and personalized support:
- Assistance with study design
- RNA purification
- Library preparation and sequencing
- Pooling and sequencing
- Bioinformatics analysis
GenoScreen's assets
- Full-service provision from study design to bioinformatics analysis of the results.
- Optimized protocols for maximizing sensitivity, reliability, and reproducibility.
- Precise summary reports, detailing the most significant differences.
The high performance level of NGS technologies is largely based on bioinformatics methods capable of reassembling entire sequences (contigs or scaffolds) from a very large number of fragments of genomes or transcriptomes.
The first task is to organize and assemble these fragments of information on a reference genome. The goal is to identify variants with the standard, then, to understand their roles and impacts (annotation and interpretation). To this end, our bioinformaticians use semi-automated processing algorithms.
This requires considerable calculation and analytical capacities, knowing genomics is now one of the leading consumers of high-performance calculation (HPC) and big data services.
Along with the calculation power, the data storage constitutes another major challenge in terms of volume, performance and security in genomics. In fact, GenoScreen keeps a high level of sensitivity and confidentiality of your data.
Applications - Assembly
The assembly allows the reconstitution of a whole genome using new sequencing techniques, either based on a reference genome or via a de novo approach.
This reconstruction gives a comprehensive, detailed image of genomic data that facilitates the latter’s interpretation.
GenoScreen - Assembly services provided
In order to obtain the best results, GenoScreen combines the most suitable methods for each case :
- De novo assembly, which assembles the reads generated by sequencing (of a genome or transcriptome) into longer sequences (contigs or scaffolds) with the objective of determining the entire sequence of a chromosome or a transcript.
- Mapping which aligns the short fragments on a reference sequence in FASTA format.
Our teams use several algorithms to optimize the length of the assembled contigs and thus reduce the assembly error rate as much as possible.
GenoScreen's assets
A low error rate, credits to our exclusive de novo assembly techniques.
Genomic annotations are aimed to associate the sequencing of a genome or transcriptome with exploitable biological information. By using high-quality assemblies, GenoScreen delivers the precise annotation required for understanding an organism’s biological functions.
Applications of genomic annotations
By linking sequencing data to previously acquired genomic knowledge, annotation deepens our understanding of a genome.
The main goal is, firstly, to identify variants (with the standard) and, secondly, to understand their roles and impacts. Annotation provides an understanding of an organism’s evolution or gene expression (diseases, agrifood production, etc.)
GenoScreen - Genomic annotations services provided
- Structural annotation localizes a sequence’s coding regions in order to predict the genes’ positions (open reading frames, ORFs).
- Functional annotation uses alignment techniques to compare the sequences with previously annotated databases. It can also be used to screen for sequence motifs that are specific for certain biological functions.
GenoScreen's assets
- Exhaustive searching within functional genomics reference databases (UniProt / GeneOntology, etc.)
- Significant high-performance computing capacity.
- Expertise in big data techniques.
- In-depth knowledge of microbial functional genomics.
News
-
New EFSA Guidance for microorganisms (2025) : What changes and how GenoScreen help you stay fully compliant
Following its agenda to regulate the Food and Feed industry with up-to-date regulatory texts, the EFSAsignificantly raised the technical expectations for the characterisation and risk assessment of microorganisms used in food and feed. As of November 4th, 2025, dossiers that rely on outdated genomic...
-
How to scientifically and regulatory validate a probiotic or postbiotic Strain (EFSA, FDA)?
What are the requirements for marketing a probiotic or postbiotic strain? Developing supplements or functional foods containing a biotic strain involves much more than simply demonstrating their effectiveness. This process is essential for obtaining regulatory approvals from authorities such as the...
-
From prebiotic to LBP: Measuring functional effects in the gut microbiome
An imbalanced microbiome is not only characterized by a lack of diversity or abundance of species. What truly matters are the functions expressed:








